
The experiment was split longitudinally into 7 separate runs at roughly 32 samples per run.Ī typical session (comprised sample preparation and staining) took me 7 hours, starting from getting the samples out of cryopreservation, counting them (for normalizing the input stain size), aliquoting cells, staining itself, and fixation.Īcquisition was performed on a full-spectrum 4-lasers Cytek Aurora, which gave me sweet advantages over traditional flow cytometers: (A) no compensation control setup at each run (shaving off 40 mins per run) (B) flexibility in panel design (because it is a full spectrum cytometer) and (C) high flow rate with low abort rate, i.e. Well, analysis is always an on-going process anyway, only stopped when there is enough to convince reviewer #2. The experiment was completed successfully and analysis is currently underway. This is very typical of a common PhD trajectory as a candidate progresses: problems mostly coming from trying to scale things up because time is running out. This was where things get difficult, not in term of science but in term of logistics. My population of interest, rare antigen-specific memory B cells (defined as live CD3 –CD19 +CD27 +Probe +), usually in the neighborhood of 100 events per million events in healthy donors 1. Usually, a flow cytometry experiment poses no problem, but particularly for this project I had to query for rare events.  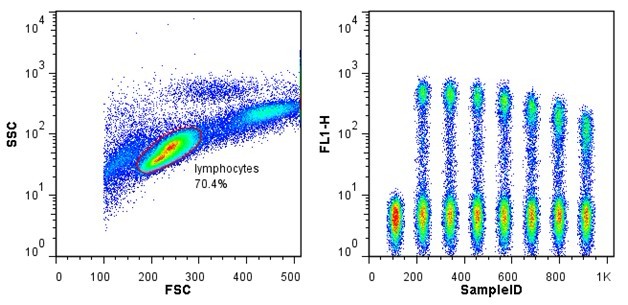
Recently I had to process ~200 human PBMC samples by flow cytometry for an immunophenotyping project.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
